PEtab data format specification
Format version: 1
This document explains the PEtab data format.
Purpose
Providing a standardized way for specifying parameter estimation problems in systems biology, especially for the case of Ordinary Differential Equation (ODE) models.
Scope
The scope of PEtab is the full specification of parameter estimation problems in typical systems biology applications. In our experience, a typical setup of data-based modeling starts either with (i) the model of a biological system that is to be calibrated, or with (ii) experimental data that are to be integrated and analyzed using a computational model. Measurements are linked to the biological model by an observation and noise model. Often, measurements are taken after some perturbations have been applied, which are modeled as derivations from a generic model (Figure 1A). Therefore, one goal was to specify such a setup in the least redundant way. Furthermore, we wanted to establish an intuitive, modular, machine- and human-readable and -writable format that makes use of existing standards.
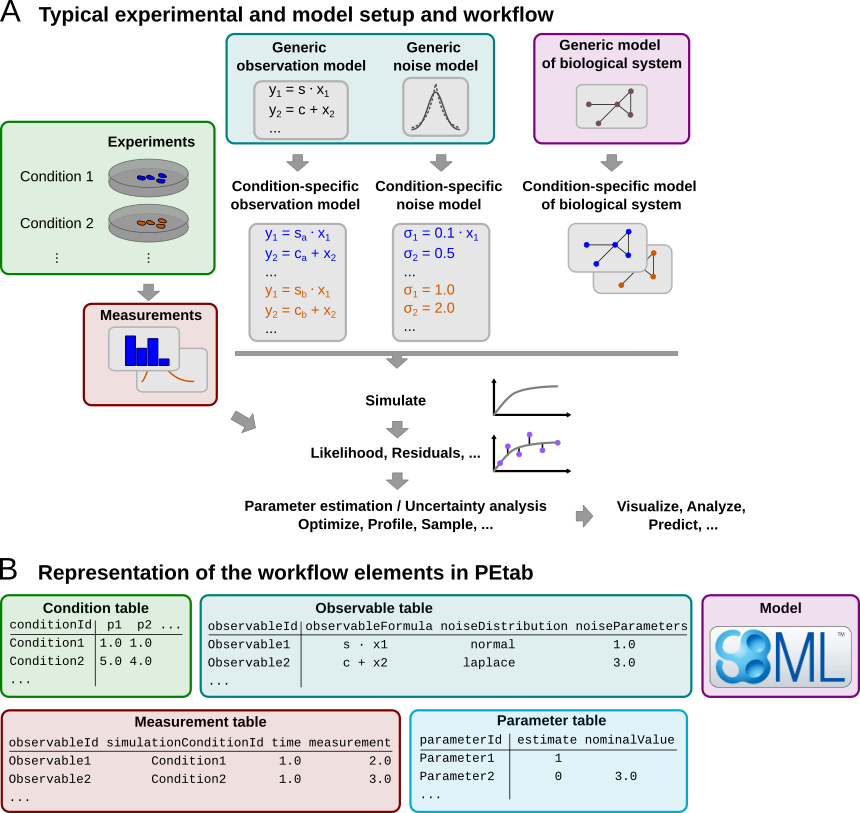
Figure 1: A common setup for data-based modeling studies and its representation in PEtab.
Overview
The PEtab data format specifies a parameter estimation problem using a number of text-based files (Systems Biology Markup Language (SBML) and Tab-Separated Values (TSV)) (Figure 2), i.e.
An SBML model [SBML]
A measurement file to fit the model to [TSV]
A condition file specifying model inputs and condition-specific parameters [TSV]
An observable file specifying the observation model [TSV]
A parameter file specifying optimization parameters and related information [TSV]
(optional) A simulation file, which has the same format as the measurement file, but contains model simulations [TSV]
(optional) A visualization file, which contains specifications how the data and/or simulations should be plotted by the visualization routines [TSV]
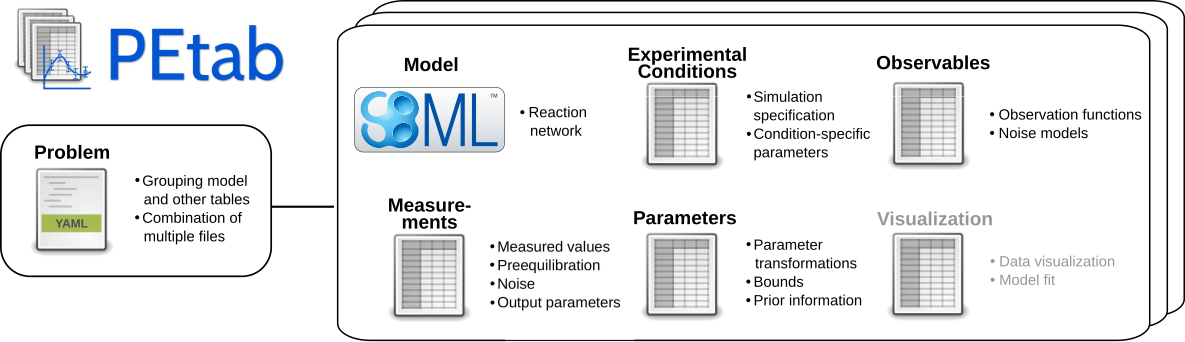
Figure 2: Files constituting a PEtab problem.
Figure 1B shows how those files relate to a common setup for data-based modeling studies.
The following sections will describe the minimum requirements of those components in the core standard, which should provide all information for defining the parameter estimation problem.
Extensions of this format (e.g. additional columns in the measurement table) are possible and intended. However, while those columns may provide extra information for example for plotting, downstream analysis, or for more efficient parameter estimation, they should not affect the optimization problem as such.
General remarks
All model entities, column names and row names are case-sensitive
Fields in “[]” are optional and may be left empty.
SBML model definition
The model must be specified as valid SBML. There are no further restrictions.
Condition table
The condition table specifies parameters, or initial values of species and compartments for specific simulation conditions (generally corresponding to different experimental conditions).
This is specified as a tab-separated value file in the following way:
conditionId |
[conditionName] |
parameterOrSpeciesOrCompartmentId1 |
… |
parameterOrSpeciesOrCompartmentId${n} |
|---|---|---|---|---|
STRING |
[STRING] |
NUMERIC|STRING |
… |
NUMERIC|STRING |
e.g. |
||||
conditionId1 |
[conditionName1] |
0.42 |
… |
parameterId |
conditionId2 |
… |
… |
… |
… |
… |
… |
… |
… |
Row- and column-ordering are arbitrary, although specifying conditionId
first may improve human readability.
Additional columns are not allowed.
Detailed field description
conditionId[STRING, NOT NULL]Unique identifier for the simulation/experimental condition, to be referenced by the measurement table described below. Must consist only of upper and lower case letters, digits and underscores, and must not start with a digit.
conditionName[STRING, OPTIONAL]Condition names are arbitrary strings to describe the given condition. They may be used for reporting or visualization.
${parameterOrSpeciesOrCompartmentId1}Further columns may be global parameter IDs, IDs of species or compartments as defined in the SBML model. Only one column is allowed per ID. Values for these condition parameters may be provided either as numeric values, or as IDs defined in the SBML model, the parameter table or both.
${parameterId}The values will override any parameter values specified in the model.
${speciesId}If a species ID is provided, it is interpreted as the initial condition of that species (as amount if hasOnlySubstanceUnits is set to True for the respective species, as concentration otherwise) and will override the initial condition given in the SBML model or given by a preequilibration condition. If
NaNis provided for a condition, the result of the preequilibration (or initial condition from the SBML model, if no preequilibration is defined) is used.${compartmentId}If a compartment ID is provided, it is interpreted as the initial compartment size.
Measurement table
A tab-separated values files containing all measurements to be used for model training or validation.
Expected to have the following named columns in any (but preferably this) order:
observableId |
[preequilibrationConditionId] |
simulationConditionId |
measurement |
time |
|---|---|---|---|---|
observableId |
[conditionId] |
conditionId |
NUMERIC |
NUMERIC|inf |
… |
… |
… |
… |
… |
(wrapped for readability)
… |
[observableParameters] |
[noiseParameters] |
|---|---|---|
… |
[parameterId|NUMERIC[;parameterId|NUMERIC][…]] |
[parameterId|NUMERIC[;parameterId|NUMERIC][…]] |
… |
… |
… |
Additional (non-standard) columns may be added. If the additional plotting functionality of PEtab should be used, such columns could be
… |
[datasetId] |
[replicateId] |
|---|---|---|
… |
[datasetId] |
[replicateId] |
… |
… |
… |
where datasetId is a necessary column to use particular plotting
functionality, and replicateId is optional, which can be used to group
replicates and plot error bars.
Detailed field description
observableId[STRING, NOT NULL, REFERENCES(observables.observableID)]Observable ID as defined in the observables table described below.
preequilibrationConditionId[STRING OR NULL, REFERENCES(conditionsTable.conditionID), OPTIONAL]The
conditionIdto be used for preequilibration. E.g. for drug treatments, the model would be preequilibrated with the no-drug condition. Empty for no preequilibration.simulationConditionId[STRING, NOT NULL, REFERENCES(conditionsTable.conditionID)]conditionIdas provided in the condition table, specifying the condition-specific parameters used for simulation.measurement[NUMERIC, NOT NULL]The measured value in the same units/scale as the model output.
time[NUMERIC OR STRING, NOT NULL]Time point of the measurement in the time unit specified in the SBML model, numeric value or
inf(lower-case) for steady-state measurements.observableParameters[NUMERIC, STRING OR NULL, OPTIONAL]This field allows overriding or introducing condition-specific versions of output parameters defined in the observation model. The model can define observables (see below) containing place-holder parameters which can be replaced by condition-specific dynamic or constant parameters. Placeholder parameters must be named
observableParameter${n}_${observableId}withnranging from 1 (not 0) to the number of placeholders for the given observable, without gaps. If the observable specified underobservableIdcontains no placeholders, this field must be empty. If it containsn > 0placeholders, this field must holdnsemicolon-separated numeric values or parameter names. No trailing semicolon must be added.Different lines for the same
observableIdmay specify different parameters. This may be used to account for condition-specific or batch-specific parameters. This will translate into an extended optimization parameter vector.All placeholders defined in the observation model must be overwritten here. If there are no placeholders used, this column may be omitted.
noiseParameters[NUMERIC, STRING OR NULL, OPTIONAL]The measurement standard deviation or
NaNif the corresponding sigma is a model parameter.Numeric values or parameter names are allowed. Same rules apply as for
observableParametersin the previous point.datasetId[STRING, OPTIONAL]The datasetId is used to group certain measurements to datasets. This is typically the case for data points which belong to the same observable, the same simulation and preequilibration condition, the same noise model, the same observable transformation and the same observable parameters. This grouping makes it possible to use the plotting routines which are provided in the PEtab repository.
replicateId[STRING, OPTIONAL]The replicateId can be used to discern replicates with the same
datasetId, which is helpful for plotting e.g. error bars.
Observables table
Parameter estimation requires linking experimental observations to the model of interest. Therefore, one needs to define observables (model outputs) and respective noise models, which represent the measurement process. Since parameter estimation is beyond the scope of SBML, there exists no standard way to specify observables (model outputs) and respective noise models. Therefore, in PEtab observables are specified in a separate table as described in the following. This allows for a clear separation of the observation model and the underlying dynamic model, which allows, in most cases, to reuse any existing SBML model without modifications.
The observable table has the following columns:
observableId |
[observableName] |
observableFormula |
|---|---|---|
STRING |
[STRING] |
STRING |
e.g. |
||
relativeTotalProtein1 |
Relative abundance of Protein1 |
observableParameter1_relativeTotalProtein1 * (protein1 + phospho_protein1 ) |
… |
… |
… |
(wrapped for readability)
… |
[observableTransformation] |
noiseFormula |
[noiseDistribution] |
|---|---|---|---|
… |
[lin(default)|log|log10] |
STRING|NUMBER |
[laplace|normal] |
… |
e.g. |
||
… |
lin |
noiseParameter1_relativeTotalProtein1 |
normal |
… |
… |
… |
… |
Detailed field description
observableId[STRING]Unique identifier for the given observable. Must consist only of upper and lower case letters, digits and underscores, and must not start with a digit. This is referenced by the
observableIdcolumn in the measurement table.[
observableName] [STRING, OPTIONAL]Name of the observable. Only used for output, not for identification.
observableFormula[STRING]Observation function as plain text formula expression. May contain any symbol defined in the SBML model (including model time
time) or parameter table. In the simplest case just an SBML species ID or anAssignmentRuletarget.May introduce new parameters of the form
observableParameter${n}_${observableId}, which are overridden byobservableParametersin the measurement table (see description there).
observableTransformation[STRING, OPTIONAL]Transformation of the observable and measurement for computing the objective function. Must be one of
lin,logorlog10. Defaults tolin. The measurements and model outputs are both assumed to be provided in linear space.
noiseFormula[NUMERIC|STRING]Measurement noise can be specified as a numerical value which will default to a Gaussian noise model if not specified differently in
noiseDistributionwith standard deviation as provided here. In this case, the same standard deviation is assumed for all measurements for the given observable.Alternatively, some formula expression can be provided to specify more complex noise models. A noise model which accounts for relative and absolute contributions could, e.g., be defined as:
noiseParameter1_observable_pErk + noiseParameter2_observable_pErk*pErk
with
noiseParameter1_observable_pErkdenoting the absolute andnoiseParameter2_observable_pErkthe relative contribution for the observableobservable_pErkcorresponding to speciespErk. IDs of noise parameters that need to have different values for different measurements have the structure:noiseParameter${indexOfNoiseParameter}_${observableId}to facilitate automatic recognition. The specific values or parameters are assigned in thenoiseParametersfield of the measurement table (see above). Any parameters namednoiseParameter${1..n}_${observableId}must be overwritten in the measurement table.Noise formulae can also contain observable parameter overrides, which are described under
observableFormulain this table. An example is when an observable formula contains an override, and a proportional noise model is used, which means the observable formula also appears in the noise formula.
noiseDistribution[STRING: ‘normal’ or ‘laplace’, OPTIONAL]Assumed noise distribution for the given measurement. Only normally or Laplace distributed noise is currently allowed (log-normal and log-Laplace are obtained by setting
observableTransformationtolog, similarly forlog10). Defaults tonormal. Ifnormal, the specifiednoiseParameterswill be interpreted as standard deviation (not variance). IfLaplaceist specified, the specifiednoiseParameterwill be interpreted as the scale, or diversity, parameter.
Noise distributions
For noiseDistribution, normal and laplace are supported. For observableTransformation, lin, log and log10 are supported. Denote by \(y\) the simulation, \(m\) the measurement, and \(\sigma\) the standard deviation of a normal, or the scale parameter of a laplace model, as given via the noiseFormula field. Then we have the following effective noise distributions.
Normal distribution:
\[\pi(m|y,\sigma) = \frac{1}{\sqrt{2\pi}\sigma}\exp\left(-\frac{(m-y)^2}{2\sigma^2}\right)\]Log-normal distribution (i.e. log(m) is normally distributed):
\[\pi(m|y,\sigma) = \frac{1}{\sqrt{2\pi}\sigma m}\exp\left(-\frac{(\log m - \log y)^2}{2\sigma^2}\right)\]Log10-normal distribution (i.e. log10(m) is normally distributed):
\[\pi(m|y,\sigma) = \frac{1}{\sqrt{2\pi}\sigma m \log(10)}\exp\left(-\frac{(\log_{10} m - \log_{10} y)^2}{2\sigma^2}\right)\]Laplace distribution:
\[\pi(m|y,\sigma) = \frac{1}{2\sigma}\exp\left(-\frac{|m-y|}{\sigma}\right)\]Log-Laplace distribution (i.e. log(m) is Laplace distributed):
\[\pi(m|y,\sigma) = \frac{1}{2\sigma m}\exp\left(-\frac{|\log m - \log y|}{\sigma}\right)\]Log10-Laplace distribution (i.e. log10(m) is Laplace distributed):
\[\pi(m|y,\sigma) = \frac{1}{2\sigma m \log(10)}\exp\left(-\frac{|\log_{10} m - \log_{10} y|}{\sigma}\right)\]
The distributions above are for a single data point. For a collection \(D=\{m_i\}_i\) of data points and corresponding simulations \(Y=\{y_i\}_i\) and noise parameters \(\Sigma=\{\sigma_i\}_i\), the current specification assumes independence, i.e. the full distributions is
Parameter table
A tab-separated value text file containing information on model parameters.
This table must include the following parameters:
Named parameter overrides introduced in the conditions table, unless defined in the SBML model
Named parameter overrides introduced in the measurement table
and must not include:
Placeholder parameters (see
observableParametersandnoiseParametersabove)Parameters included as column names in the condition table
Parameters that are AssignmentRule targets in the SBML model
SBML local parameters
it may include:
Any SBML model parameter that was not excluded above
Named parameter overrides introduced in the conditions table
One row per parameter with arbitrary order of rows and columns:
parameterId |
[parameterName] |
parameterScale |
lowerBound |
upperBound |
nominalValue |
estimate |
… |
|---|---|---|---|---|---|---|---|
STRING |
[STRING] |
log10|lin|log |
NUMERIC |
NUMERIC |
NUMERIC |
0|1 |
… |
… |
… |
… |
… |
… |
… |
… |
… |
(wrapped for readability)
… |
[initializationPriorType] |
[initializationPriorParameters] |
[objectivePriorType] |
[objectivePriorParameters] |
|---|---|---|---|---|
… |
see below |
see below |
see below |
see below |
… |
… |
… |
… |
… |
Additional columns may be added.
Detailed field description
parameterId[STRING, NOT NULL]The
parameterIdof the parameter described in this row. This has to match the ID of a parameter specified in the SBML model, a parameter introduced as override in the condition table, or a parameter occurring in theobservableParametersornoiseParameterscolumn of the measurement table (see above).parameterName[STRING, OPTIONAL]Parameter name to be used e.g. for plotting etc. Can be chosen freely. May or may not coincide with the SBML parameter name.
parameterScale[lin|log|log10]Scale of the parameter to be used during parameter estimation.
lowerBound[NUMERIC]Lower bound of the parameter used for optimization. Optional, if
estimate==0. Must be provided in linear space, independent ofparameterScale.upperBound[NUMERIC]Upper bound of the parameter used for optimization. Optional, if
estimate==0. Must be provided in linear space, independent ofparameterScale.nominalValue[NUMERIC]Some parameter value to be used if the parameter is not subject to estimation (see
estimatebelow). Must be provided in linear space, independent ofparameterScale. Optional, unlessestimate==0.estimate[BOOL 0|1]1 or 0, depending on, if the parameter is estimated (1) or set to a fixed value(0) (see
nominalValue).initializationPriorType[STRING, OPTIONAL]Prior types used for sampling of initial points for optimization. Sampled points are clipped to lie inside the parameter boundaries specified by
lowerBoundandupperBound. Defaults toparameterScaleUniform.Possible prior types are:
uniform: flat prior on linear parameters
normal: Gaussian prior on linear parameters
laplace: Laplace prior on linear parameters
logNormal: exponentiated Gaussian prior on linear parameters
logLaplace: exponentiated Laplace prior on linear parameters
parameterScaleUniform (default): Flat prior on original parameter scale (equivalent to “no prior”)
parameterScaleNormal: Gaussian prior on original parameter scale
parameterScaleLaplace: Laplace prior on original parameter scale
initializationPriorParameters[STRING, OPTIONAL]Prior parameters used for sampling of initial points for optimization, separated by a semicolon. Defaults to
lowerBound;upperBound. The parameters are expected to be in linear scale except for theparameterScalepriors, where the prior parameters are expected to be in parameter scale.So far, only numeric values will be supported, no parameter names. Parameters for the different prior types are:
uniform: lower bound; upper bound
normal: mean; standard deviation (not variance)
laplace: location; scale
logNormal: parameters of corresp. normal distribution (see: normal)
logLaplace: parameters of corresp. Laplace distribution (see: laplace)
parameterScaleUniform: lower bound; upper bound
parameterScaleNormal: mean; standard deviation (not variance)
parameterScaleLaplace: location; scale
objectivePriorType[STRING, OPTIONAL]Prior types used for the objective function during optimization or sampling. For possible values, see
initializationPriorType.objectivePriorParameters[STRING, OPTIONAL]Prior parameters used for the objective function during optimization. For more detailed documentation, see
initializationPriorParameters.
Visualization table
A tab-separated value file containing the specification of the visualization
routines which come with the PEtab repository. Plots are in general
collections of different datasets as specified using their datasetId (if
provided) inside the measurement table.
Expected to have the following columns in any (but preferably this) order:
plotId |
[plotName] |
[plotTypeSimulation] |
[plotTypeData] |
|---|---|---|---|
STRING |
[STRING] |
[LinePlot(default)|BarPlot|ScatterPlot] |
[MeanAndSD(default)|MeanAndSEM|replicate;provided] |
… |
… |
… |
… |
(wrapped for readability)
… |
[datasetId] |
[xValues] |
[xOffset] |
[xLabel] |
[xScale] |
|---|---|---|---|---|---|
… |
[datasetId] |
[time(default)|parameterOrStateId] |
[NUMERIC] |
[STRING] |
[lin|log|log10|order] |
… |
… |
… |
… |
… |
… |
(wrapped for readability)
… |
[yValues] |
[yOffset] |
[yLabel] |
[yScale] |
[legendEntry] |
|---|---|---|---|---|---|
… |
[observableId] |
[NUMERIC] |
[STRING] |
[lin|log|log10] |
[STRING] |
… |
… |
… |
… |
… |
… |
Detailed field description
plotId[STRING, NOT NULL]An ID which corresponds to a specific plot. All datasets with the same plotId will be plotted into the same axes object.
plotName[STRING, OPTIONAL]A name for the specific plot.
plotTypeSimulation[STRING, OPTIONAL]The type of the corresponding plot, can be
LinePlot,BarPlotandScatterPlot. Default isLinePlot.plotTypeData[STRING, OPTIONAL]The type how replicates should be handled, can be
MeanAndSD,MeanAndSEM,replicate(for plotting all replicates separately), orprovided(if numeric values for the noise level are provided in the measurement table). Default isMeanAndSD.datasetId[STRING, NOT NULL, REFERENCES(measurementTable.datasetId), OPTIONAL]The datasets which should be grouped into one plot.
xValues[STRING, OPTIONAL]The independent variable, which will be plotted on the x-axis. Can be
time(default, for time resolved data), or it can beparameterOrStateIdfor dose-response plots. The corresponding numeric values will be shown on the x-axis.xOffset[NUMERIC, OPTIONAL]Possible data-offsets for the independent variable (default is
0).xLabel[STRING, OPTIONAL]Label for the x-axis. Defaults to the entry in
xValues.xScale[STRING, OPTIONAL]Scale of the independent variable, can be
lin,log,log10ororder. Theordervalue should be used if values of the independent variable are ordinal. This value can only be used in combination withLinePlotvalue for theplotTypeSimulationcolumn. In this case, points on x axis will be placed equidistantly from each other. Default islin.yValues[observableId, REFERENCES(measurementTable.observableId), OPTIONAL]The observable which should be plotted on the y-axis.
yOffset[NUMERIC, OPTIONAL]Possible data-offsets for the observable (default is
0).yLabel[STRING, OPTIONAL]Label for the y-axis. Defaults to the entry in
yValues.yScale[STRING, OPTIONAL]Scale of the observable, can be
lin,log, orlog10. Default islin.legendEntry[STRING, OPTIONAL]The name that should be displayed for the corresponding dataset in the legend and which defaults to the value in
datasetId.
Extensions
Additional columns, such as Color, etc. may be specified.
Examples
Examples of the visualization table can be found in the Benchmark model collection, for example in the Chen_MSB2009 model.
YAML file for grouping files
To link the SBML model, measurement table, condition table, etc. in an unambiguous way, we use a YAML file.
This file also allows specifying a PEtab version (as the format is not unlikely to change in the future).
Furthermore, this can be used to describe parameter estimation problems comprising multiple models (more details below).
The format is described in the schema ../petab/petab_schema.yaml, which allows for easy validation.
Parameter estimation problems combining multiple models
Parameter estimation problems can comprise multiple models. For now, PEtab allows to specify multiple SBML models with corresponding condition and measurement tables, and one joint parameter table. This means that the parameter namespace is global. Therefore, parameters with the same ID in different models will be considered identical.